CURRENT PROJECTS
The Research Alliance Leibniz INFECTIONS continues to focus on antimicrobial resistance (AMR). The World Health Organisation (WHO) describes the spread of AMR as one of the greatest public health threats. A multidisciplinary and interdisciplinary approach is required in order to tackle the problem in a results-oriented and strategically forward-looking manner. Resistant microorganisms not only arise and spread as a result of the selection pressure of antimicrobial therapies, but are also linked to human activities. These include:
- impacts on the environment
- the type of agriculture
- public health systems
- and the economic and social conditions that determine how medicines are developed, sold and used.
In order to understand why drug resistance develops and how it spreads, experts from many disciplines – not just medicine – need to work closely together. Accordingly, Leibniz INFECTIONS draws on the expertise of 16 Leibniz Institutes and other scientific partners, including the Robert Koch Institute (RKI) and the Friedrich Loeffler Institute (FLI).
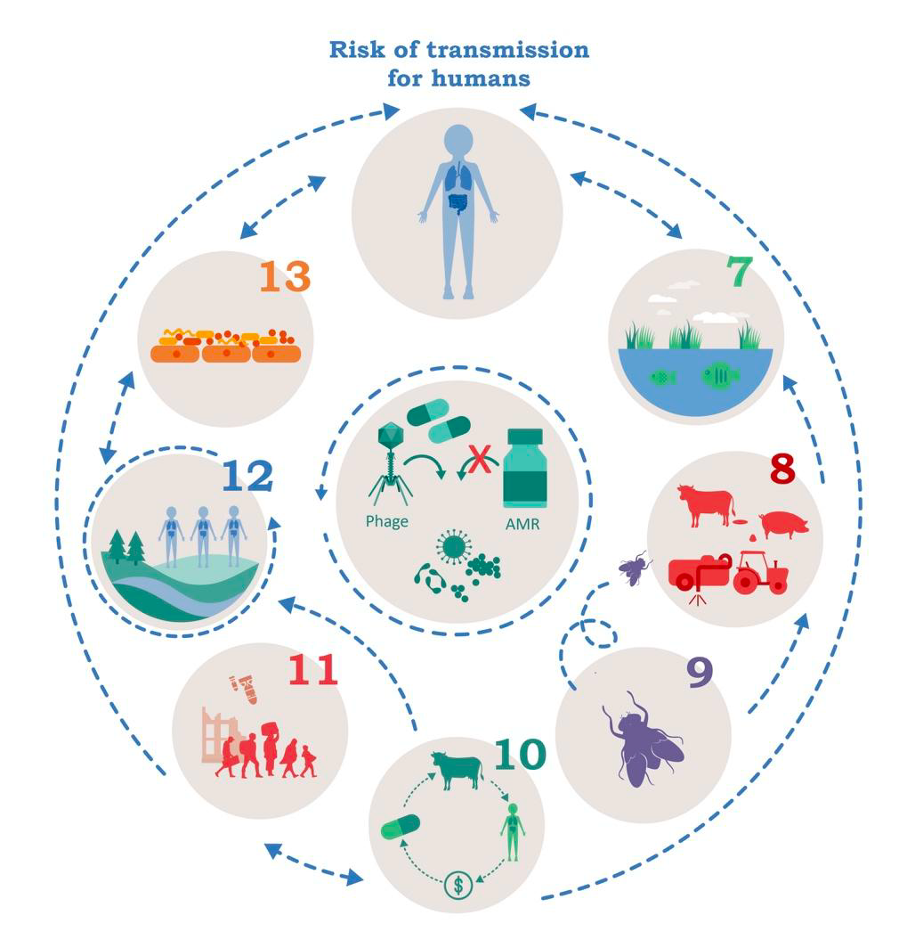
Seven research projects have been designed bringing together various partners. The interdisciplinary project teams (IPTs) are investigating the transmission of AMR from various sources – namely human, environmental, agricultural and socio-economic. The aim is to identify the general drivers of AMR, assess the risk of transmission to human health and develop counter strategies. The IPTs are numbered consecutively from 7 to 13 to distinguish them from IPTs 1 to 6 from the previous funding period (see figure). Further information can be found below in the project descriptions.
Water is vital for human life, but also plays a key role as a vector and reservoir for AMR transmission across urban and rural areas. Our results suggest that AMR abundance and diversity is higher in urban waters and sediments compared to rural water bodies. Wastewater treatment plants (WWTPs) are a major source of AMR in urban water sources. In addition, rural lakes sediments and water derived from farmlands are major sources of environmental AMR.
We will assess the risk of human contact with AMR through environmental water by leveraging our accumulated data from INFECTIONS in the previous funding period and performing bioinformatic comparison with sequences of clinical isolates in our collections. Analyses will be extended to fungal pathogens, including mobile genetic elements. Future water collection will comprise waste water sampled in partner countries where antibiotics / antifungals are used in agriculture. The phage content of the samples will be determined. We are investigating the phageome (all bacteriophages in the water) and whether they might provide promising alternatives to control AMR carrying bacteria. In pilot studies, we will attempt to remove AMR containing bacteria from wastewater using environmentally detected phage in laboratory based studies.
Participating Institutes: ATB, BNITM, DSMZ, FZB, HKI, IGB, IZW
With steadily rising of global meat consumption and the growing number of livestock animals, antibiotic resistances (AMR) spread by livestock manure is an increasingly problem that affects animals, humans, and environmental health. Because manure is used as fertilizer for agricultural fields in the sense of a sustainable circular bioeconomy, there is a potential source of AMR transmission into the human food chain. In the interdisciplinary project team 8 (IPT8), we focus on a One Health approach to track the dynamics of microbial communities and fly larvae in livestock slurry tanks used for agricultural field fertilization. Here, different farm and laboratory slurry storage conditions will be studied with a focus on the persistence of AMR bacteria including foodborne pathogens such as Salmonella, ESBL-producing E. coli and non-tuberculous mycobacteria. We aim to monitor the abundance of AMR and microbiome changes in farm and laboratory slurry tanks. The influence of seasonal temperature changes, slurry type and tank filling levels will be investigated. We will detect and quantify AMR pathogens in slurry with microbiological cultivation. To assess the microbiome diversity in slurry, 16S rDNA sequencing will be performed. The occurrence and uptake of AMR bacteria from slurry by flies will be investigated. The pathogen load of flies and their larvae will be determined by bacteriological and molecular techniques. The results from this research will enable us to deduce practical and effective measures to reduce AMR bacteria in livestock manure by identifying the optimal manure management conditions.
Participating Institutes: ATB, BNITM, DSMZ, FZB, IGB, IZW, ZALF
The interdisciplinary Project Team 9 (IPT9) investigates how flies can spread antimicrobial-resistant (AMR) pathogens from livestock to humans and assesses the resulting health risks. The study focuses on the common housefly (Musca domestica), but also includes other fly species found near livestock. We will conduct field experiments to determine fly species, population densities, movement patterns, and pathogen loads in both Germany and Ghana. Laboratory studies will examine how flies ingest, retain, and shed AMR pathogens like multidrug-resistant enterobacteria, vancomycin-resistant enterococci, non-tuberculous mycobacteria, and Candida species. Using ecological data and experimental findings, models will be developed to predict fly-driven pathogen spread and transmission and evaluate the risk to humans. The team will also assess how environmental factors affect transmission dynamics and test the effectiveness of various control strategies. Multiple Leibniz institutes contribute their expertise in entomology, microbiology, ecology, and risk analysis to deliver a comprehensive understanding of fly-mediated AMR spread.
Participating Institutes: ATB, BNITM, DSMZ, FLI, FZB, HKI, IOER, ZALF
The unregulated and on a massive scale use of antibiotics in livestock farming is a major contributor to the global spread of antimicrobial resistance (AMR). In many low-income settings, lack of veterinary oversight and weak regulatory systems result in inappropriate antibiotic use. IPT10 aims to develop and test scalable interventions to promote responsible antibiotic use in resource-constrained environments.
We combine three strategies: (1) behavior change communication campaigns targeting livestock keepers, (2) the use of affordable molecular diagnostic tools for animal health, and (3) digital platforms for monitoring antibiotic usage and resistance data. The project is implemented in collaboration with partners in Burundi, Rwanda, Tanzania, Ghana, and the Democratic Republic of Congo.
IPT10 builds on ongoing projects: AMRAfrica (a digital monitoring tool), ADA (novel diagnostics), and mobile AMR laboratories operated by the East African Community. These initiatives provide the foundation for piloting and evaluating integrated AMR control strategies. The evidence will support policy development and guide effective stewardship practices in settings where veterinary services are limited.
Participating Institutes: IfW, BNITM, FZB, RKI
Policy makers often assume some countries with certain time-invariant characteristics are generally – or per se – better at preventing antimicrobial resistance (AMR) than others. IPT11 explores an alternative hypothesis that builds on recent evidence suggesting significant localized heterogeneity in AMR dynamics at the level of drug-disease or drug-pathogen pairings – both within and across countries.
Our main focus is on tuberculosis. For each relevant antibiotic drug-pathogen pairing, we combine data from a range of public sources, including the Institute for Health Metrics and Evaluation (IHME), to identify countries that for a given pairing contribute (as drivers) to the global burden of disease / mortality more than similar countries in terms of epidemiology, demography and socioeconomic conditions. The goal is to identify antibiotic access barriers that may e.g. prevent the local use of first-best evidence-based drug combinations in an afflicted country. To study the impact of acute crises, such as war, evidence from Ukraine will be used – a country with high burdens of AMR bacterial infections, such as MDR TB and MRSA. To approach the dilemma between investment incentives and profit maximization versus global access to effective antibiotics when these are developed in the private sector, the strategic design of optimal incentives in line with evidence-based treatment guidelines and the influence of international organizations promoting the introduction and prudent use of antibiotics in the global market are also modeled.
Participating Institutes: Kiel Institute, BNITM, FZB, RKI
Non-tuberculous mycobacteria (NTM) are a diverse group of bacteria commonly found in the environment, particularly in water and soil. They are increasingly causing infections in humans, particularly in individuals with weak immune systems or pre-existing lung conditions such as cystic fibrosis, and are also known to infect a variety of animal species. These infections are often difficult to treat because many NTM are naturally resistant to multiple antibiotics.
It is thought that people get infected with NTM mainly through contact with environmental sources that contain these bacteria. However, which environmental sources currently pose the highest risk to susceptible individuals remains unknown.
This project aims to collect and analyze NTM from different environmental sources like soil and water, as well as from plant and animals across Germany. The bacteria will be examined to determine how common they are, how genetically diverse they are and whether they are resistant to the antibiotics that are normally used to treat NTM disease. These environmental NTM will also be compared to clinical isolates from human patients using modern molecular tools.
The results will help us better understand potential sources and transmission routes of NTM, evaluate infection risks, and contribute to the development of new strategies for prevention and treatment. Promising new therapies, including bacteriophages and novel compounds, will be tested on these strains in collaboration with project IPT13.
Participating Institutes: ATB, DSMZ, FZB, FLI, IGB, IZW, NTUS, RKI, UoS
In IPT13, the complex interactions of polymicrobial species will be investigated further. Once again, lung epithelial air-liquid-interface cells will be used as the model surface to culture the biofilm. As the next stage lung organoids will be established to provide a more complex growth system. Three groups of microorganisms will be studied: bacteria – M. abscessus, S. maltophilia, fungi: C. albicans, C. auris, A. fumigatus, and virus: influenza and SARS-CoV.
The aims of this project will be to assess the interactions of these microbial species by live cell microscopy, analyse how each species responds to each other in a mixed biofilm via proteomics, and explore phage isolates and novel antibiotics against the polymicrobial biofilms for therapeutic control.

